Tools portal
The CSDMS software engineers create tools to help CSDMS members couple and run models.
BMI Builder
The BMI Builder helps developers build a Basic Model Interface for their model by generating template source code which is then customized by the user for their particular model. Users provide some basic metadata for their model (such as names of input and output items, units, etc.) to the bmi builder which then creates creates BMI implementation files in their language of choice. The user then go through these file of files and implements the labeled sections.
Links
- Source code on GitHub
- Documentation on how to use the BMI Builder is provided on GitHub in the read.me file
BMI Tester
The BMI Tester is a Python package that provides command-line utilities for testing Python classes that implement the Basic Model Interface (BMI). BMI Tester also provides a Python interface to the tester that allows users to run tests programmatically. The package is easily extendable so that new tests can be added to the suite.
Links
Standard Names registry
The Standard Names registry is a Python library for working with CSDMS Standard Names.
The CSDMS Standard Names provide a comprehensive set of naming rules and patterns that are not specific to any particular modeling domain. They were also designed to have many other nice features such as parsability and natural alphabetical grouping. CSDMS Standard Names always consist of an object part and a quantity/attribute part and the quantity part may also have an operation prefix which can consist of multiple operations.
standard_names is a Python package to define, query, operate, and manipulate CSDMS Standard Names. It is distributed with a comprehensive list of the currently defined Standard Names.
The standard_names package consists of two basic classes:
- StandardName: a single name
- NamesRegistry: a collection of names.
The standard_names package has a complete set of unit tests that are continually tested on Mac, Linux, and Windows. It is distributed both as source code and as Anaconda packages from Anaconda Cloud.
Links
- Documentation for the standard_names package
- Source code
- The current Standard Names Registry on GitHub
WMT
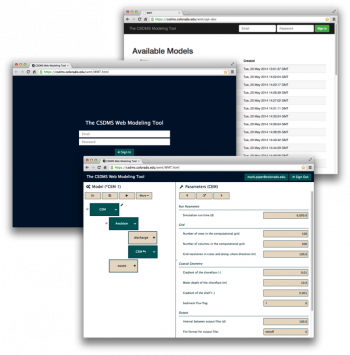
The CSDMS Web Modeling Tool, WMT, is a web application that allows users on a desktop, laptop or tablet computer to build and run standalone or coupled Earth surface dynamics models on a supercomputer.
With WMT, users can:
- Design a model from a set of CSDMS components
- Edit model parameters
- Save models to a web-accessible server
- Share saved models with the community
- Submit model runs to an HPC system
- Download simulation results
The following components are currently available in WMT.
To see how you can use WMT to configure and run a standalone or a coupled model on the web, check out the WMT tutorial, or jump right into using WMT at https://csdms.colorado.edu/wmt.
For more information on WMT, including links to its documentation and its source code repositories, please see the CSDMS Web Modeling Tool wiki page.
Dakotathon
Dakota is a software toolkit developed at Sandia National Laboratories that provides an interface between models and a library of analysis methods, including support for sensitivity analysis, uncertainty quantification, optimization, and calibration techniques. Dakota is a powerful tool, but its learning curve is steep: the user not only must understand the structure and syntax of the Dakota input file, but also must develop intermediate code that allows Dakota to set up and run a model, read outputs from the model, and calculate a response statistic from the outputs.
The CSDMS Dakota Interface, or Dakotathon, is a Python package that wraps and extends Dakota’s file-based user interface. It simplifies the process of configuring and running a Dakota experiment, and it allows a Dakota experiment to be scripted. Dakotathon creates the Dakota input file and provides a generic analysis driver. Any model componentized in the CSDMS modeling framework automatically works with Dakotathon. Dakotathon has a plugin architecture, so models not wrapped into the CSDMS modeling framework can be accessed by Dakotathon by programmatically extending a template; an example is provided in the Dakotathon distribution.
For more information on Dakotathon, including links to its documentation and its source code repository, please see the Dakotathon wiki page.
Landlab
LANDLAB is an open-source Python library of code that enables users to easily build unique earth surface processes models to address specific hypotheses. The Landlab library contains gridding engines for building regular and irregular grids, process components that act on grid variables, tools for storing and sharing data among the grid and components, and plotting and analysis tools. Process components represent individual processes occurring at the earth’s surface (e.g. growth and death of vegetation, surface water infiltration, linear diffusion of sediment), or drivers of these processes (e.g. rainfall, solar radiation). Users can build models by combining existing process components, or build new components if a required process component does not exist. Landlab was developed by members of the CSDMS community with support from CSDMS and the National Science Foundation (two ACI - SI2 grants).
Why build another modeling tool?
Landlab has the rare distinction of being built to make modeling accessible to a wide variety of earth scientists, including those who do not traditionally use computational models. In contrast, most other models are designed to answer a specific hypothesis posed by the developers.
What does this mean for new users?
- Landlab is extensively documented.
- Users can contact developers with issues.
- There are tutorials to teach users how to use Landlab.
- There are classroom tutorials to teach students about different surface processes.
- Many Landlab tutorials are accessible through Hydroshare, an online collaborative environment for sharing data and models, which allows users to test-out Landlab without installing it locally.
- Landlab has integrated testing to ensure that new code contributions will not break existing code.
- There is documentation to make contributing code a bit easier.
Landlab Components and Models
Numerous people have contributed components and models to landlab. Of these, 53 are described in the CSDMS repository and are listed below.
| Program | Description | Developer | Download | PyMT |
|---|---|---|---|---|
| ChannelProfiler | The ChannelProfiler extracts and plots channel networks from a landlab grid. | Barnhart, Katy | ||
| ChiFinder | Calculate Chi Indices | Hobley, Daniel | ||
| DepressionFinderAndRouter | Find depressions on a topographic surface. | Hobley, Dan | ||
DepthDependentDiffuser
|
Soil depth-dependent linear hillslope diffuser | Glade, Rachel | ||
| DepthDependentTaylorDiffuser | This component implements a depth-dependent Taylor series diffusion rule, combining concepts of Ganti et al. (2012) and Johnstone and Hilley (2014). | Glade, Rachel | ||
| DetachmentLtdErosion | Simulate detachment limited sediment transport. | Adams, Jordan | ||
Drainage Density
|
Component for calculating drainage density in Landlab given a channel network | Shobe, Charles | ||
ErosionDeposition
|
Landlab component for fluvial erosion/deposition. | Shobe, Charles | ||
| ExponentialWeatherer | Exponential soil production function in the style of Ahnert (1976) | Glade, Rachel | ||
| FastscapeEroder | Compute fluvial erosion using stream power theory (“fastscape” algorithm) | Hobley, Daniel | ||
| FireGenerator | This component generates a random fire event or fire time series from the Weibull statistical distribution. | Adams, Jordan | ||
| Flexure | Deform the lithosphere with 1D or 2D flexure. | Hutton, Eric | ||
| FlowAccumulator | Component to accumulate flow and calculate drainage area. | Barnhart, Katy | ||
| FlowDirectorD8 | Single-path (steepest direction) flow direction with diagonals on rasters. | Barnhart, Katy | ||
| FlowDirectorDinf | Flow direction on a raster grid by the D infinity method. | Barnhart, Katy | ||
| FlowDirectorMFD | Multiple-path flow direction with or without out diagonals. | Barnhart, Katy | ||
| FlowDirectorSteepest | Single-path (steepest direction) flow direction without diagonals. | Barnhart, Katy | ||
| FractureGridGenerator | Create a 2D grid with randomly generated fractures. | Tucker, Greg | ||
GrainHill
|
Cellular automaton model of hillslope evolution | Tucker, Gregory | ||
| GroundwaterDupuitPercolator | The GroundwaterDupuitPercolator solves the Boussinesq equation for flow in an unconfined aquifer over an impermeable aquifer base and calculates groundwater return flow to the surface. | Litwin, David | ||
| HackCalculator | Calculate Hack parameters. | Barnhart, Katy | ||
HyLands
|
The HyLands model simulates the impact of bedrock landslides on topographic evolution and sediment dynamics. | Campforts, Benjamin | ||
| LakeMapperBarnes | Temporarily fills depressions and reroutes flow across them | Hobley, Daniel | ||
| Landlab | Python software framework for writing, assembling, and running 2D numerical models | Tucker, Greg | ||
| Landslides | Landlab component that simulates landslide probability of failure as well as mean relative wetness and probability of saturation. | Strauch, Ronda | ||
| LateralEroder | Laterally erode neighbor node through fluvial erosion. | Langston, Abigail | ||
| LinearDiffuser | Landlab component that models soil creep as a linear diffusion process | Tucker, Greg | ||
| Lithology | Create a Lithology object with different properties | Barnhart, Katy | ||
| LossyFlowAccumulator | Component to calculate drainage area and accumulate flow, while permitting dynamic loss or gain of flow downstream. | Hobley, Dan | ||
| NormalFault | NormalFault implements relative rock motion due to a normal fault. | Barnhart, Katy | ||
OverlandFlow
|
Component simulating overland flow using a 2-D numerical approximation of the shallow-water equations following the de Almeida et al., 2012 algorithm for storage-cell inundation modeling. | Adams, Jordan | ||
| OverlandFlowBates | This component simulates overland flow using the 2-D numerical model of shallow-water flow over topography using the Bates et al. (2010) algorithm for storage-cell inundation modeling. | Adams, Jordan | ||
| PerronNLDiffuse | Nonlinear diffusion, following Perron (2011). | Hobley, Daniel | ||
| PotentialEvapotranspiration | Calculates potential evapotranspiration | Nudurupati, Sai | ||
| PotentialityFlowRouter | Multidirectional flow routing using a novel method. | Hobley, Daniel | ||
| PrecipitationDistribution | Generate random sequence of precipitation events | Adams, Jordan | ||
| Radiation | Compute 1D and 2D total incident shortwave radiation. | Nudurupati, Sai | ||
SPACE
|
Landlab component for 2-D calculation of fluvial sediment transport and bedrock erosion | Shobe, Charles | ||
| SedDepEroder | Compute fluvial erosion using using “tools and cover” theory | Hobley, Daniel | ||
| SinkFiller | Fill sinks in a landscape to the brim, following the Barnes et al. (2014) algorithms. | Hobley, Daniel | ||
SoilInfiltrationGreenAmpt
|
Landlab component that calculates soil infiltration based on the Green-Ampt solution. | Rengers, Francis | ||
| SoilMoisture | Compute the decay of soil moisture saturation at storm-interstorm time period | Nudurupati, Sai | ||
| SpatialPrecipitationDistribution | Generate random sequence of spatially-resolved precipitation events | Hobley, Daniel | ||
| SpeciesEvolver | Evolve life in a landscape. | Lyons, Nathan | ||
| SteepnessFinder | Calculate steepness and concavity indices from gridded topography | Hobley, Daniel | ||
| StreamPowerSmoothThresholdEroder | Compute fluvial erosion using stream power theory with a numerically smoothed threshold | Tucker, Greg | ||
| TaylorNonLinearDiffuser | Model non-linear soil creep after Ganti et al. (2012) | Glade, Rachel | ||
Terrainbento
|
A Python package for multi-model analysis in long-term drainage basin evolution | Barnhart, Katy | ||
| TransportLengthHillslopeDiffuser | Transport length hillslope diffusion. | Mouchene, Margaux | ||
| VegCA | Landlab component that simulates inter-species plant competition using a 2D cellular automata model. | Nudurupati, Sai | ||
| Vegetation | Model plant dynamics using multiple representative plant species | Nudurupati, Sai | ||
| ZoneController | Controls zones and populates them with taxa. | Lyons, Nathan | ||
| ZoneTaxon | A zone-based taxon | Lyons, Nathan |
Permafrost Benchmark System
The Permafrost Benchmark System (PBS) wraps the command-line ILAMB benchmarking software with a customized version of WMT, and adds tools for uploading CMIP5-compatible model outputs and benchmark datasets. The PBS allows users to access and run ILAMB remotely, without having to install software or data locally; a web browser on a desktop, laptop, or tablet computer is all that’s needed.
For more information on the PBS, including links to documentation and source code repositories, please see the PBS home page at https://permamodel.github.io/pbs.
If you're already set up with a login on the PBS,
jump right to it at https://csdms.colorado.edu/pbs.